 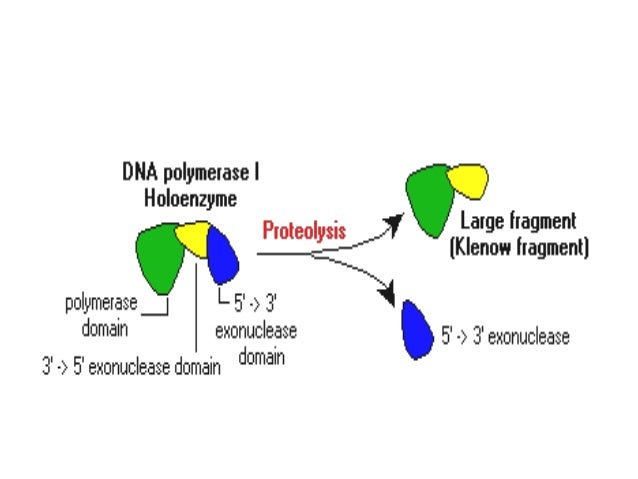
Much of modern biotechnology and molecular biology relies on the ability to manipulate a specific nucleic acid target, but in order to efficiently detect, read, or use that target sequence, it must be available at sufficient quantities in the application. The ability presented here to produce long, discrete DNA products in an isothermal reaction extends the scope of isothermal amplification to enable more useful applications of these promising methods. We demonstrate that two features of gp32 enable this amplification: a facilitation of primer strand invasion into double-stranded DNA, and a suppression of non-homologous primer annealing and nonspecific amplification. In addition to the discrete amplicon products, this method also produces higher molecular weight products consisting of multiple repeated copies of the amplicon and template DNA. In contrast to existing methods, this amplification requires only the single-stranded DNA-binding protein gp32 from bacteriophage T4 and a strand-displacing DNA polymerase. Here we report the amplification of discrete target fragments of several kilobases at 37 ☌ from both double- and single-stranded circular template DNA using specific primer pairs. Current isothermal methods result in the generation of short fragments (<150 base pairs) or highly branched long DNA products. These results suggest that the polymerase and exonuclease sites of Klenow are physically separate in solution and exhibit different substrate structural requirements for activity.Isothermal amplification methods for detection of DNA and RNA targets have expanded significantly in recent years, promising a new wave of simple and rapid molecular diagnostics. On the other hand, the polymerase activity of Klenow fragment required that the primer terminus be at least six base pairs downstream from the base with the biotin-avidin complex. In the presence of the biotin binding protein avidin, the exonucleolytic activity of Klenow fragment requires that the primer terminus be at least 15 base pairs downstream from the base with the biotin-avidin complex. A DNA duplex with a biotin covalently linked to a specific base has been prepared. Klenow fragment and T4 DNA polymerase are able to polymerize onto duplexes incapable of strand separation, whereas T7 DNA polymerase seems to require that the primer terminus be at least three bases from the cross-linked base pair. The exonucleolytic action of T4 and T7 DNA polymerases requires that only two and three bases respectively melt out for excision of nucleotides from the primer terminus. The action of Klenow fragment on these duplexes indicates that the polymerase site does not require that the DNA duplex undergo strand separation for activity, whereas the exonuclease site requires that at least four base pairs of the primer strand must melt out for the exonucleolytic removal of nucleotides from the primer terminus. This and similar duplexes are substrates for the polymerase and exonuclease activities of the Klenow fragment of Escherichia coli DNA polymerase I and T4 and T7 DNA polymerases. A DNA duplex covalently cross-linked between specific bases has been prepared. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
